An international team of researchers from industry and academia has recently published a novel computational method to find new medicinal drugs.
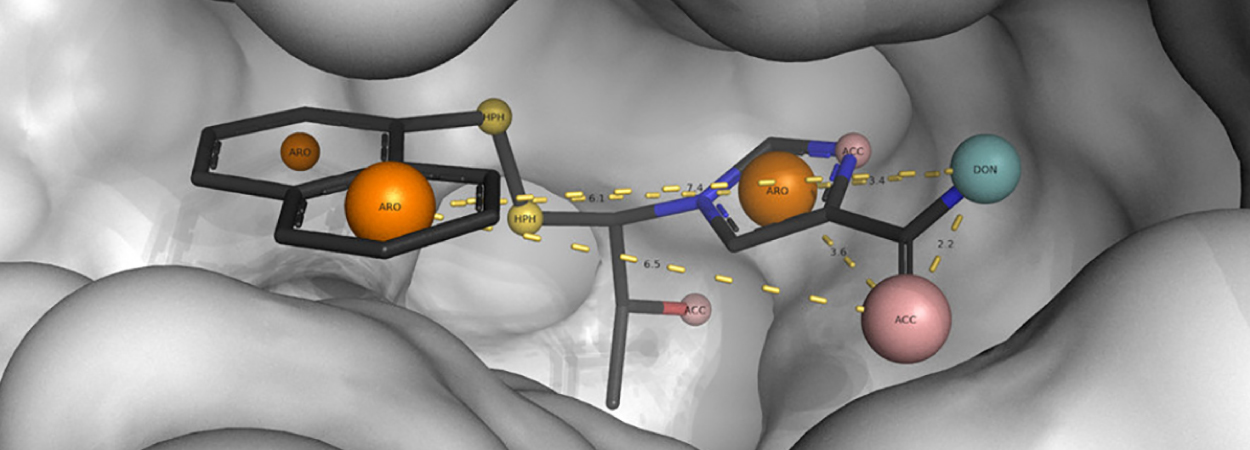
This bioinformatics method uses structures of known protein-ligand complexes to find important interaction points (called PIPs) between the two. The 3D arrangement of these points is then used to guide a search for potential drugs in a large database containing millions of molecules. The method has important repercussions in drug discovery, particularly to improve productivity in the drug discovery process.
The method has been used successfully in a commercial setting at Oxford Drug Design, a biotechnology company which focuses on the discovery of new antibiotics.
This research collaboration consists of Dr Jean-Paul Ebejer (Centre for Molecular Medicine and Biobanking, University of Malta), Prof. Charlotte Deane, Prof. Garrett Morris, Wing Ki Wong (University of Oxford), and Prof. Paul Finn (University of Buckingham and Oxford Drug Design).
The study, titled 'Ligity: A Non-Superpositional, Knowledge-Based Approach to Virtual Screening', appeared in the peer-reviewed Journal of Chemical Information and Modelling (an American Chemical Society publication). This publication is open access.
For further information about this research kindly contact Dr Jean Paul Ebejer.